Understanding Human Ribosome Biogenesis
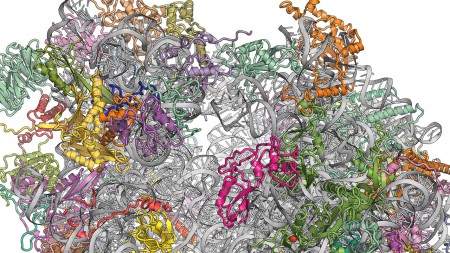
“Crystal structure of the budding yeast ribosome.” Marat Yusupov (CNRS, Strasbourg).
We are characterizing ribosome assembly pathways in human cells and budding yeast. We are particularly interested by RNA processing and RNA modification steps.
Eukaryotic ribosome biogenesis is a complex process, which is initiated in the nucleolus, proceeds into the nucleoplasm and is finalized in the cytoplasm. All three RNA polymerases are involved and tightly regulated by proto-oncogenes and tumor suppressors. In budding yeast, a reference eukaryotic model, it took >20 years to identify over 200 trans-acting assembly factors. These and several dozens small nucleolar RNAs (snoRNAs) are required to assemble ribosomes.
Recent work has highlighted that ribosome synthesis is far more complicated in human cells than anticipated from work on yeast. While the general architecture of the pre-rRNA processing pathway is evolutionarily conserved, specific individual reactions proceed differently. Unsurprisingly, human cells have evolved additional unique factors, more than 75 of which have now been uncovered (see www.RibosomeSynthesis.Com). Yeast is clearly no longer a sufficient model for studying ribosomopathies at the molecular level, and work on human cells is imperative in order to achieve true therapeutic advances.

