Molecular, Cellular & Pathophysiological Aspects of Ribosome Biogenesis

“A man reflecting on human ribosome biogenesis.” Art by Sybille Mercat.
Ribosomes are essential cellular nanomachines, converting the genetic information encoded in messenger RNAs (mRNAs) into proteins playing structural or catalytic roles in the cell. The human ribosome contains four rRNAs and eighty ribosomal proteins organized in two subunits, each carrying out specialized functions in translation. The decoding site, where the mRNA is “read”, is in the small-subunit (40S). The large-subunit (60S) harbors the peptidyl transferase center, where amino acids are joined together to form polypeptides, and the exit tunnel, from where nascent proteins emerge. At the ribosome’s functional core is rRNA, a ribozyme that catalyzes the critical steps of decoding and amino acid polymerization.
Ribosome synthesis is initiated in the nucleolus, a specialized compartment of the nucleus. The nucleolus is a long-standing disease biomarker and a recently demonstrated target for cancer therapy. There are many connections between nucleolar structure/function and the regulation of the expression of the anti-tumor protein p53.
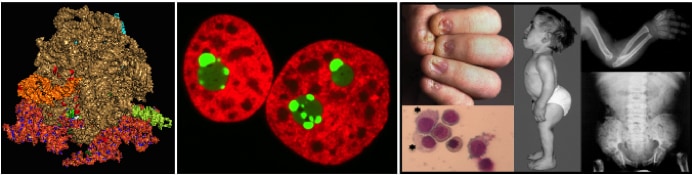
Ribosomes are indispensable to gene expression and disruption of their synthesis leads inevitably to severe disease. Ribosomopathies are human syndromes caused by ribosome assembly dysfunction resulting from mutations in ribosomal proteins or ribosome assembly factors. Ribosomopathies are characterized by a triad of symptoms: hematological defects, skeletal problems, and increased cancer susceptibility. Notable examples include: Diamond blackfan anemia and X-linked Dyskeratosis congenita.