Understanding Human Nucleolar Structure
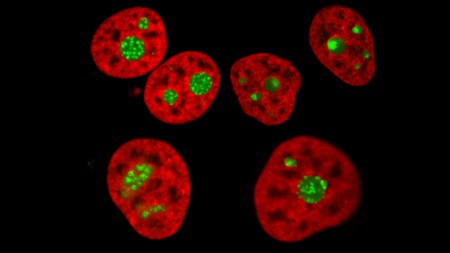
“A human cell line expressing histone H2B in red and a nucleolar antigen in green imaged by spin-disc confocal microscopy in normal (top left) and perturbed conditions.” (Lafontaine Lab).
We are characterizing the principles of nucleolar structure integrity maintenance and dynamics.
Ribosome synthesis is initiated in the nucleolus, a prominent highly dynamic nuclear compartment central to gene expression. The morphology of the nucleolus is indicative of its function and cell health, and as such it is a potent disease biomarker and a recently demonstrated target for cancer therapy. In interphase, human nucleoli display three morphologically distinct subcompartments. During mitosis, nucleoli undergo a cycle of drastic disassembly-reassembly that parallels other mitotic events.
We have developed a powerful software for characterizing nucleolar disruption qualitatively and quantitatively with a precise numerical index: the iNo score (index of nucleolar disruption, Nicolas et al. 2016). For this, we have implemented a high-resolution, high-throughput screening platform with visual readout (an automated confocal spindisc microscope). As proof-of-concept, we depleted human cells expressing a fluorescent nucleolar reporter construct of each of the eighty ribosomal proteins, one by one, and systematically assessed nucleolar integrity. Unexpectedly, we found only a few ribosomal proteins to be required to maintain nucleolar integrity, the strongest contributors being uL5 and uL18. These are precisely the two factors which, together with the 5S rRNA, form a trimeric ribonucleoprotein complex important in regulating the homeostasis of the anticancer protein p53 (Nicolas et al. 2016). The dataset presenting the involvement of each individual human ribosomal protein in mature rRNA accumulation, pre-rRNA processing, nucleolar structure, and p53 steady-state accumulation is available at Www.RibosomalProteins.Com.

